Ancient population genomics
In our group, we are interested in understanding how human societies have shaped ancient and present-day biodiversity, by developing novel computational tools that efficiently capture the evolutionary information encoded within the genomes of ancient, museum and modern populations. A few examples of our current interests are:
a) Molecular and evolutionary mechanisms underlying early domestication stages.
We now understand that our impact on biodiversity commenced thousands of years ago, with the domestication of plants and animals and with the often eradication of their wild relatives. Beyond when and where plants and animals were domesticated, however, it remains unclear how domestication actually took place. What molecular and evolutionary mechanisms are involved at early domestication stages? To what extent these represent preferential entry routes into domestication? Can epigenetic and microbiome changes somehow contribute to early domestication stages?
b) Conservation genomics leveraging museum specimens.
Biodiversity conservation has thus far focused on a subset of iconic species, vastly ignoring the fundamental and major importance of arthropods in environmental health, for example through pollination. As past archives, arthropod specimens stored from entomological collections represent a fantastic resource for tracking ancient diversity across space and time, and to document how industrialisation, global warming and pollution alter the arthropods' biology. Using time-stamped genomic series, we can go beyond monitoring genetic diversity, and also explore the trajectory of adaptive alleles, the deleterious load within populations (inbreeding genetic load vs. purging), as well as more fundamental shifts in life-history traits such as generation times.
c) Evolutionary models and computational tools.
Leveraging the additional temporal information provided by genomic time-series, we are developing statistical models to estimate key evolutionary parameters in conservation and domestication, such as generation time changes and dominance coefficients. While shorter generation times could confer greater adaptive potential in front of sudden environmental challenges and lifestyle transitions, dominance coefficients are crucial to accurately identify and quantify selection, including both adaptive alleles and genomic load. Ancient DNA data, however, is not devoid of challenges as fragmented and chemically damaged post-mortem. Such degradation is non-random, but shaped by the true epigenetic status of the sample. By exploiting rather than masking this damage in the DNA, we develop tools to produce lower error rates in the sequence data compared to generic methods, while additionally reconstructing ancient epigenetic information, helping understand the role of phenotypic plasticity into evolution.
Interested in joining our group? Contact me.
Librado P. 2024. Reconstructing generation intervals over time. Nature Reviews Genetics, 25(11):745-746 . DOI:10.1038/s41576-024-00766-2
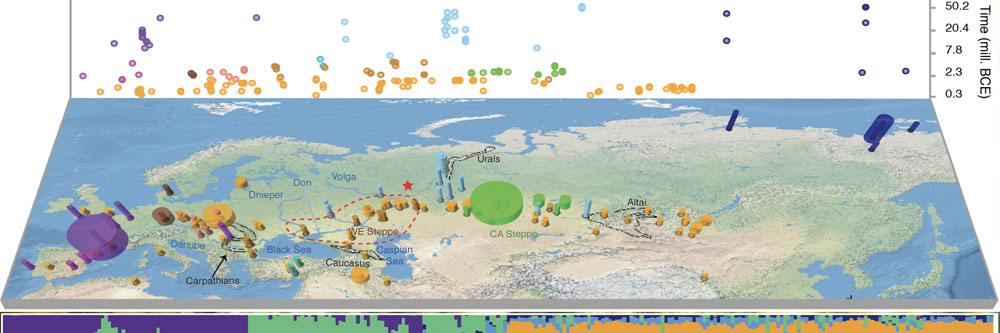
Librado P, Tressières G, Chauvey L, Fages A, Khan N, Schiavinato S, Calvière-Tonasso L, Kusliy MA, Gaunitz C, Liu X, Wagner S, Der Sarkissian C, Seguin-Orlando A, Perdereau A, Aury JM, Southon J, Shapiro B, Bouchez O, Donnadieu C, Collin YRH, Gregersen KM, Jessen MD, Christensen K, Claudi-Hansen L, Pruvost M, Pucher E, Vulic H, Novak M, Rimpf A, Turk P, Reiter S, Brem G, Schwall C, Barrey É, Robert C, Degueurce C, Horwitz LK, Klassen L, Rasmussen U, Kveiborg J, Johannsen NN, Makowiecki D, Makarowicz P, Szeliga M, Ilchyshyn V, Rud V, Romaniszyn J, Mullin VE, Verdugo M, Bradley DG, Cardoso JL, Valente MJ, Antunes MT, Ameen C, Thomas R, Ludwig A, Marzullo M, Prato O, Gianni GB, Tecchiati U, Granado J, Schlumbaum A, Deschler-Erb S, Mráz MS, Boulbes N, Gardeisen A, Mayer C, Döhle HJ, Vicze M, Kosintsev PA, Kyselý R, Peške L, O'Connor T, Ananyevskaya E, Shevnina I, Logvin A, Kovalev AA, Iderkhangai TO, Sablin MV, Dashkovskiy PK, Graphodatsky AS, Merts I, Merts V, Kasparov AK, Pitulko VV, Onar V, Öztan A, Arbuckle BS, McColl H, Renaud G, Khaskhanov R, Demidenko S, Kadieva A, Atabiev B, Sundqvist M, Lindgren G, López-Cachero FJ, Albizuri S, Trbojević Vukičević T, Rapan Papeša A, Burić M, Rajić Šikanjić P, Weinstock J, Vilaró DA, Codina F, Dalmau CG, de Llorens JM, Pou J, de Prado G, Sanmartí J, Kallala N, Torres JR, Maraoui-Telmini B, Belarte Franco MC, Valenzuela-Lamas S, Zazzo A, Lepetz S, Duchesne S, Alexeev A, Bayarsaikhan J, Houle JL, Bayarkhuu N, Turbat T, Crubézy É, Shingiray I, Mashkour M, Berezina NY, Korobov DS, Belinskiy A, Kalmykov A, Demoule JP, Reinhold S, Hansen S, Wallner B, Roslyakova N, Kuznetsov PF, Tishkin AA, Wincker P, Kanne K, Outram A, Orlando L. 2024. Widespread horse-based mobility arose around 2,200 BCE in Eurasia. Nature, 631(8022):819-825. DOI:10.1038/s41586-024-07597-5
Barquera R, Del Castillo-Chávez O, Nägele K, Pérez-Ramallo P, Hernández-Zaragoza DI, Szolek A, Rohrlach AB, Librado P, Childebayeva A, Bianco RA, Penman BS, Acuña-Alonzo V, Lucas M, Lara-Riegos JC, Moo-Mezeta ME, Torres-Romero JC, Roberts P, Kohlbacher O, Warinner C, Krause J. 2024. Ancient genomes reveal insights into ritual life at Chichén Itzá. Nature, 630(March 2023). DOI:10.1038/s41586-024-07509-7